Serious lupus: Strong genetic risk factor on its path
Last Updated on August 21, 2022 by Joseph Gut – thasso
August 21, 2022 – Systemic lupus erythematosus (SLE) is a common, incurable autoimmune disease that affects millions of individuals worldwide, with a particularly high prevalence among women. A genetic variant, called HLA-DRB1*03:01, is the greatest risk factor for the condition, which involves inflammation in many vital organs, and can lead to severe disability and death.
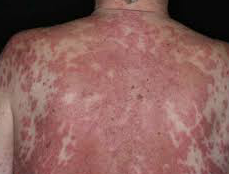 In a study recently published in Communications Biology, investigators found that a protein coded by that HLA variant triggers a cascade of molecular and cellular effects that can cause the inflammatory symptoms seen in lupus patients.
In a study recently published in Communications Biology, investigators found that a protein coded by that HLA variant triggers a cascade of molecular and cellular effects that can cause the inflammatory symptoms seen in lupus patients.
For the first time, researchers found the enigmatic mechanism that genetically predisposes people to the worst effects of the most typical forms of lupus. This type of findings could potentially facilitate the discovery of safe, simple and effective treatments for SLE by targeting this new pathway. The results support a novel theory how genetic variants of the kind of HLA-DRB1*03:01 can lead to autoimmune diseases independent of antigen presentation, the traditionally studied mechanism of autoimmunity which has been long proposed but, so far, not directly proven.
 Overall, the researchers identified a chain of events in cell culture, as well as a mouse model of the disease, that demonstrate how the abnormalities that can cause lupus develop from the first effect of the risk gene, to signaling, all the way to immune abnormalities and clinical manifestations of lupus. Thus, they identified the HLA-DRB1*03:01 allele as a major genetic risk factor in systemic lupus erythematosus (SLE). They showed that in the presence of interferon gamma (IFN-γ), a short DRB1*03:01-encoded allelic epitope (termed “lupus epitope (LE)”) activates a characteristic lupus transcriptome in mouse and human macrophages. This epitope triggers a cascade of SLE-associated cellular aberrations, including endoplasmic reticulum stress, unfolded protein response, mitochondrial dysfunction, necroptotic cell death, and production of pro-inflammatory cytokines, i.e., a whole pathway leading to a phenotypic lupus endpoint.
Overall, the researchers identified a chain of events in cell culture, as well as a mouse model of the disease, that demonstrate how the abnormalities that can cause lupus develop from the first effect of the risk gene, to signaling, all the way to immune abnormalities and clinical manifestations of lupus. Thus, they identified the HLA-DRB1*03:01 allele as a major genetic risk factor in systemic lupus erythematosus (SLE). They showed that in the presence of interferon gamma (IFN-γ), a short DRB1*03:01-encoded allelic epitope (termed “lupus epitope (LE)”) activates a characteristic lupus transcriptome in mouse and human macrophages. This epitope triggers a cascade of SLE-associated cellular aberrations, including endoplasmic reticulum stress, unfolded protein response, mitochondrial dysfunction, necroptotic cell death, and production of pro-inflammatory cytokines, i.e., a whole pathway leading to a phenotypic lupus endpoint.
The findings of this study are reminiscent of previous findings in rheumatoid arthritis, another HLA-associated disease, that have paved the way for the development of small molecules to effectively treat arthritis in mice, used as an animal model of the human disease. Human trials in RA with those compounds are being carried out, and there is hope that the novel findings will lead to similar efforts to ease the burden of millions of lupus patients as well.
On the other hand, we need to understand that the human leukocyte antigen complex (HLA-complex) is an immensely complicated complex of genes on chromosome 6 in humans which encode cell-surface proteins responsible for the regulation of the immune system. HLA genes are highly polymorphic, which means that they have many different alleles, allowing them to fine-tune the adaptive immune system, but leave them with a enormous potential of interaction on different pathways. People with certain HLA antigens are more likely to develop certain autoimmune diseases, such as type I diabetes, ankylosing spondylitis, rheumatoid arthritis, celiac disease, myasthenia gravis, inclusion body myositis, Sjögren syndrome, and narcolepsy, apart from SLE discussed here. Furthermore, some patients carrying particular alleles out of the HLA complex may suffer from serious, if not fatal drug-induced adverse reactions (SADRs) such as Stevens-Johnson Syndrome (SJS) or Toxic Epidermal Necrolysis (TEN) such as Asian patients carrying HLA-B*15:02 or HLA-B*5801 alleles (see the post by thasso on the subject). Thus, it remains very interestingly to see, if at the end of the day, a very specific treatment (molecule) will successfully be developed , very specifically addressing just one HLA allele (i.e.,HLA-DRB1*03:01) without any side effects on related pathways and without any adverse effects on related HLA allele carriers.
See here a sequence on the complexity of SLE and possibly underlying genetics:


Leave a Reply
You must be logged in to post a comment.
Optional: Social Subscribe/Login