Genetics behind some serious adverse drug reactions (SADRs)

Last Updated on December 28, 2020 by Joseph Gut – thasso
December 28, 2020 – Some 95% of people have gene variants that may negatively affect their response to at least one drug, leading to unexpected or adverse drug reactions (ADRs) or, if unlucky, to a serious, if not fatal, adverse drug reaction (SADR). In effect, SADRs are a major cause of morbidity and mortality worldwide, amounting to an overall incidence of SADRs in hospitalized patients in the US at an estimated 6.2–6.7%, with an incidence of fatal SADRs is estimated to be 0.15–0.3%, with studies in Europe and Australia yielding similar estimates.
 Some SADRs may be predictable, based upon a drug’s pharmacodynamic and pharmacokinetic properties. Many, however, appear to be idiosyncratic (i.e., seemingly not explicable rationally and thusly reflecting our lack of understanding their underlying mechanisms). Genetic factors may underlie susceptibility to SADRs and the identification of predisposing genotypes may improve patient management through the prospective selection of appropriate (drug) candidates. Several specific SADRs may emphasize the role of predisposing genetic risk factors on the development of SADRs. Among those are examples such as i) severe drug-induced cutaneous reactions, ii) drug-induced long QT and Torsades de pointes, iii) statin-induced myotoxicity, and iv) drug-induced liver injury, to name just a few.
Some SADRs may be predictable, based upon a drug’s pharmacodynamic and pharmacokinetic properties. Many, however, appear to be idiosyncratic (i.e., seemingly not explicable rationally and thusly reflecting our lack of understanding their underlying mechanisms). Genetic factors may underlie susceptibility to SADRs and the identification of predisposing genotypes may improve patient management through the prospective selection of appropriate (drug) candidates. Several specific SADRs may emphasize the role of predisposing genetic risk factors on the development of SADRs. Among those are examples such as i) severe drug-induced cutaneous reactions, ii) drug-induced long QT and Torsades de pointes, iii) statin-induced myotoxicity, and iv) drug-induced liver injury, to name just a few.
Severe drug-induced cutaneous reactions
Thus, very serious cutaneous reactions include Stevens Johnson Syndrome (SJS) and Toxic Epidermal Necrosis (TEN) which occur with very high prevalence, for example, in Han Chinese patients who are under anticonvulsant carbamazepine (CBZ) therapy and are carriers of the HLA-B*1502 allelic gene variant. In fact, SJS and TEN develop with very many medications (see here a list of such medications) and are strongly associated with a selection of different gene allelic variants throughout other ethnic groups.
Drug-induced long QT and Torsades de Pointes
For drug-induced long QT syndrome (LQTS) and Torsades de pointes (TdP) multiple clinical risk markers, notably hypokalaemia, hypomagnesaemia, underlying bradycardia, coexistent heart disease and female gender, have been identified and are considered in the analysis of any comprehensive genotype-phenotype evaluation. Ten disease genes for congenital LQTS have been identified. The two most commonly affected genes (KCNQ1 and KCNH2 (HERG)) encode the voltage-gated potassium channels underlying the currents IKs and IKr, respectively. Six of these genes also encode ion channel proteins or ancillary (function-modifying) subunits, and the remaining two disease genes, ANK2 and CAV3, do not encode ion channels but are thought to modify channel 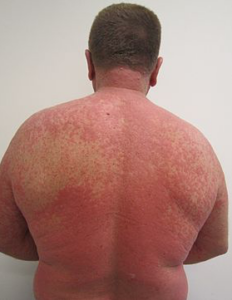 function. The ankyrin-B protein encoded by the ANK2 gene appears to target calcium-handling proteins to appropriate membrane subdomains within myocytes, whereas the CAV3 gene encodes a caveolar protein that appears to modify sodium channel function. Each disease-associated mutation above upsets the balance between inward depolarizing and outward repolarizing currents during cardiac repolarization in favour of increased net inward current, resulting in prolonged action potentials and hence, increased QT intervals on the surface ECG. Virtually all drugs that cause torsades de pointes block IKr/human ether-a-go-go-related (HERG) channels. QT prolongation is an established adverse effect of anti-arrhythmic medicines, but can also be caused by a wide range of non-cardiac medicines, including antibiotics, antihistamines, opioid analgesics and complementary medicines. There is a database available which lists drugs that are implicated in LOTS and TdP based on their interactions with the genes (i.e., their coded for proteins) discussed above.
function. The ankyrin-B protein encoded by the ANK2 gene appears to target calcium-handling proteins to appropriate membrane subdomains within myocytes, whereas the CAV3 gene encodes a caveolar protein that appears to modify sodium channel function. Each disease-associated mutation above upsets the balance between inward depolarizing and outward repolarizing currents during cardiac repolarization in favour of increased net inward current, resulting in prolonged action potentials and hence, increased QT intervals on the surface ECG. Virtually all drugs that cause torsades de pointes block IKr/human ether-a-go-go-related (HERG) channels. QT prolongation is an established adverse effect of anti-arrhythmic medicines, but can also be caused by a wide range of non-cardiac medicines, including antibiotics, antihistamines, opioid analgesics and complementary medicines. There is a database available which lists drugs that are implicated in LOTS and TdP based on their interactions with the genes (i.e., their coded for proteins) discussed above.
Statin-induced myotoxicity
Statin drugs (i.e., HMG CoA reductase inhibitors, which currently comprise Atorvastatin (Lipitor), Fluvastatin (Lescol), Lovastatin (Mevacor), Pravastatin (Pravachol), Rosuvastatin (Crestor), Simvastatin (Zocor), and Pitavastatin (Livalo)) reduce the incidence of primary and secondary coronary artery disease in patients at risk. Generally, statin drugs are regarded as both safe and efficacious. However, over time, muscle complications have emerged as a major ADR problem with statin drugs. Statin-induced muscle complications appear to be dose-dependent and they have been described diagnostically as myalgia (focal or diffuse muscle pain), myopathy (pain in the absence of inflammation), myositis (pain accompanied by a systemic inflammatory response) and rhabdomyolysis (severe muscle damage accompanied by damage in another organ, most notably the kidneys. Clinically seen is rhabdomyolysis the most important and most severe (even fatal in some cases) SADR associated with statin therapy. Genetically and molecularly seen, predispositions of patients for statin-induced muscle problems are certainly complex and may depend on allelic variants in genes coding for i) drug metabolizing enzymes, ii) drug transporters, and iii) for enzymes involved in lipid metabolism and homeostasis. Thus, across all statin drugs, variants of genes such as CYP2C8, CYP3A5, OATP2, OATP1B1 (SLCO1B1), OATPC, OATP1B3 (SLCO1B3), ABCB1 (MDR1), CPT 2, AMPD, PYGM, and COQ2 have been implicated in the susceptibility of patients for developing muscle problems.
Drug-induced liver injury (DILI)
Drug-induced liver injury (DILI) is a major reason for regulatory actions against marketing approval, removal from the market place and restriction of prescribing indications. Most drugs responsible for severe DILI are not predictable hepatotoxins. Rather, they are completely safe over a wide range of doses for the vast majority of treated patients, but severely toxic to a small subset of patients. The onset of liver injury is characteristically delayed weeks or months after starting therapy, and the liver injury is generally diffuse. In most instances, it is unclear what makes some individuals susceptible to liver toxicity, but available data support a substantial genetic contribution. The table below summarizes some of the studies that have reported statistically significant associations between variants in specific genes and susceptibility to DILI. These risk factors involve genetic variants in two major categories of genes: the highly polymorphic genes in the major histocompatibility locus on chromosome 6, which encode antigen-presenting proteins, and various polymorphic genes that encode drug metabolizing enzymes. Because the populations in these studies have generally been small, so far only common polymorphisms have been tested as susceptibility alleles for DILI.
Table summarising some selected reports of genetic associations with DILI
| Drug | Gene(s) | Drug class | Form of toxicity | Cases/controls | Refs |
|---|---|---|---|---|---|
| Ximelagatran | DRB1*07 | Oral thrombin inhibitor
|
Elevation in transaminase | 74/130 | 1 |
|
|
DQA1*02
|
|
|
|
|
| Tolcapone
|
UGT1A16
|
Catechol-O-methyltransferase inhibitor
|
Asymptomatic liver transaminase elevation
|
135/274
|
2
|
| Diclofenac | UGT2B7 | NSAID | Range from acute liver failure to non-specific symptoms with transaminase elevation
|
24/48 | 3 |
| CYP2C8 | |||||
|
|
ABCC2
|
|
|
|
|
| Tranilast‡
|
UGT1A1
|
TGF-α antagonist
|
Unconjugated hyper-bilirubinaemia
|
127/909
|
4
|
| Isoniazid | CYP2E1 | Antibiotic | Elevation in serum transaminases
|
49/269 | 5 |
|
|
NAT2
|
|
|
|
|
| Isoniazid | GSTM1 | Antibiotic | Icteric hepatitis (serum bilirubin > 3.0 mg/dL)
|
37/33 | 6 |
|
|
NAT2
|
|
Afterthoughts
Overall, these are not rare genetic disorders. Genetic variants are normal differences in the DNA code between different people. Effectively, they’re what make some people have blue eyes and others brown, for example. But the impact of genetic variation is not yet widely taken into account in medicine. Most prescribers think that because it is genetic, this is a very rare event. They still think it is academic. They think it’s something for the future. In fact, 95% of people have a gene variant that is known to affect their response to at least one drug. This is often due to changes in the way that it is broken down by the body. If it is metabolised slowly, even a standard dose of the drug can build up to high levels in the body and cause serious adverse reactions. Breaking the problem down to the level of the individual patient, it becomes a rather trist issue of life and death, as the following case sadly illustrates. Thus, Professor Henk-Jan Guchelaar, from the Clinical Pharmacy Department at the University of Leiden in the Netherlands, who has spent the last two decades trying to get the link between medicine and our genes, just mentions this story of one of his patients with fatal outcome as a point in case. The patient told Prof. Guchelaar about his wife, who had breast cancer and underwent surgery. The prognosis was very good. The tumour was removed by the surgeon but, to prevent micro-metastases, his wife had to receive six courses of the chemotherapy drug fluorouracil. During the second infusion of the drug, the patient collapsed, went to the intensive care unit and died. A blood sample taken later showed she was the carrier of a variant of the DPYD gene that is known to predispose patient to potentially fatal effects under fluorouracil therapy. The tragedy in this case is that the woman had a very good prognosis (i.e., the tumor was gone) and had she been tested beforehand the therapy with fluorouracil for the known potentially fatal genetic variants of DYPD, it was very likely that she would not have had this severe drug reaction, and would still be alive.
Prescribing medicines and thinking about risk-benefit ratios, also including risk analysis for the individual patients susceptibilities for SDARS might constitute a true step forward in personalised medicine, or even better in theragenomic medicine. The latter takes into account the whole of genetic backgrounds (inclusive confounding factors) in the therapy of the individual with a given drug for a given indication.
See here a sequence on genetic factors affecting adverse drug reactions (ADRs). See also some earlier articles by thasso on SJS, LOT, Rhabdomyolysis, and DILI.

